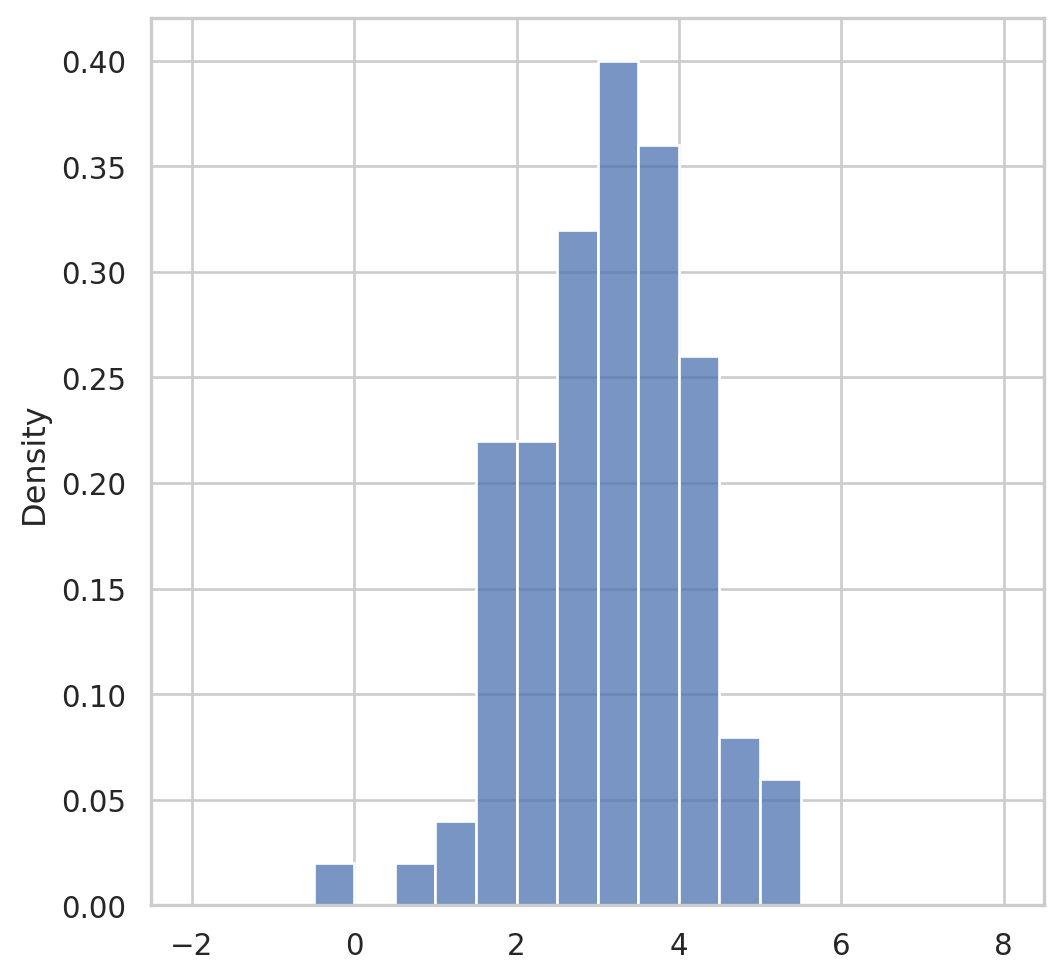
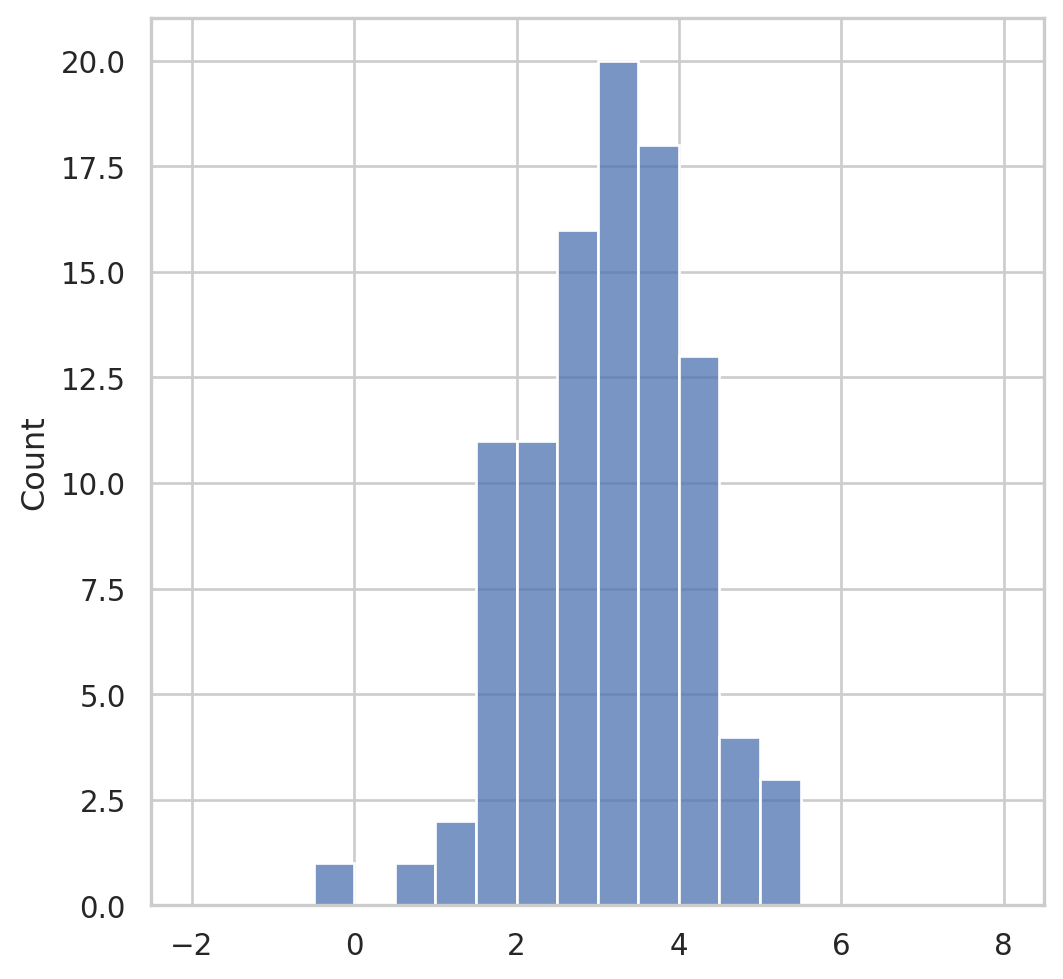
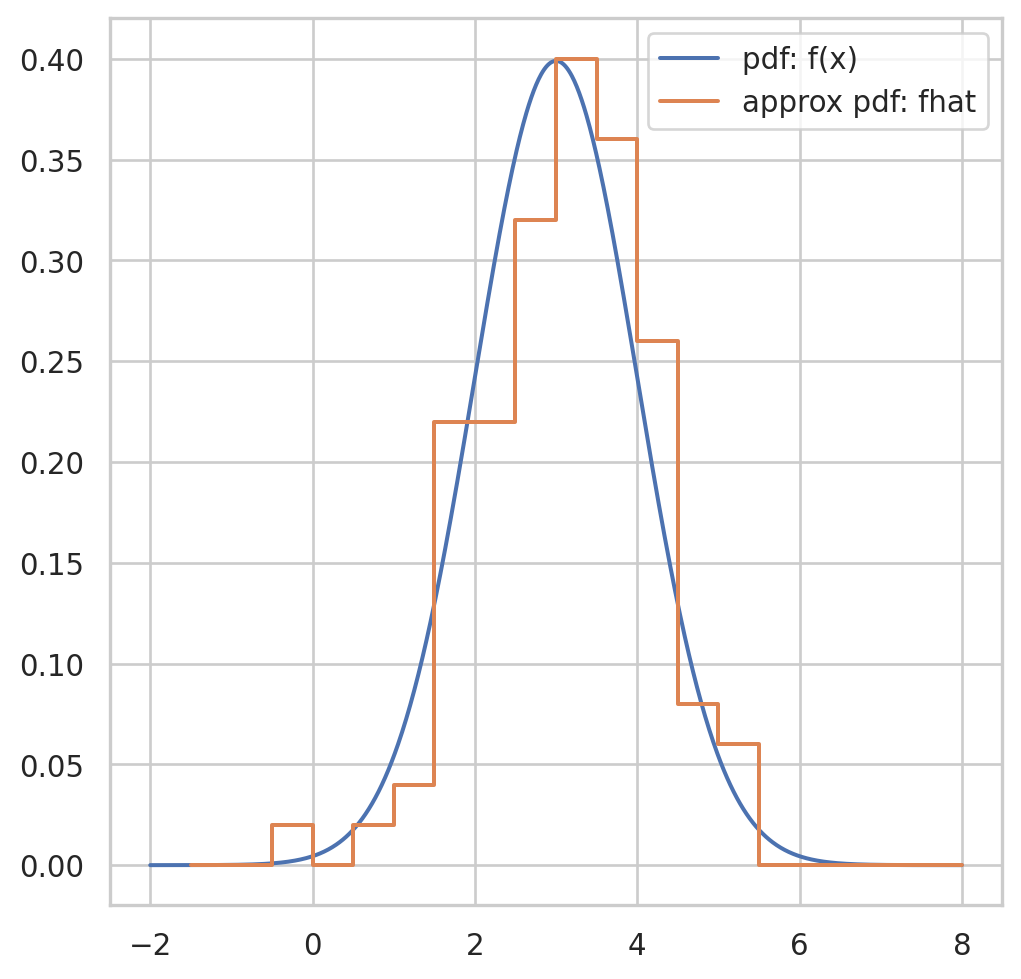
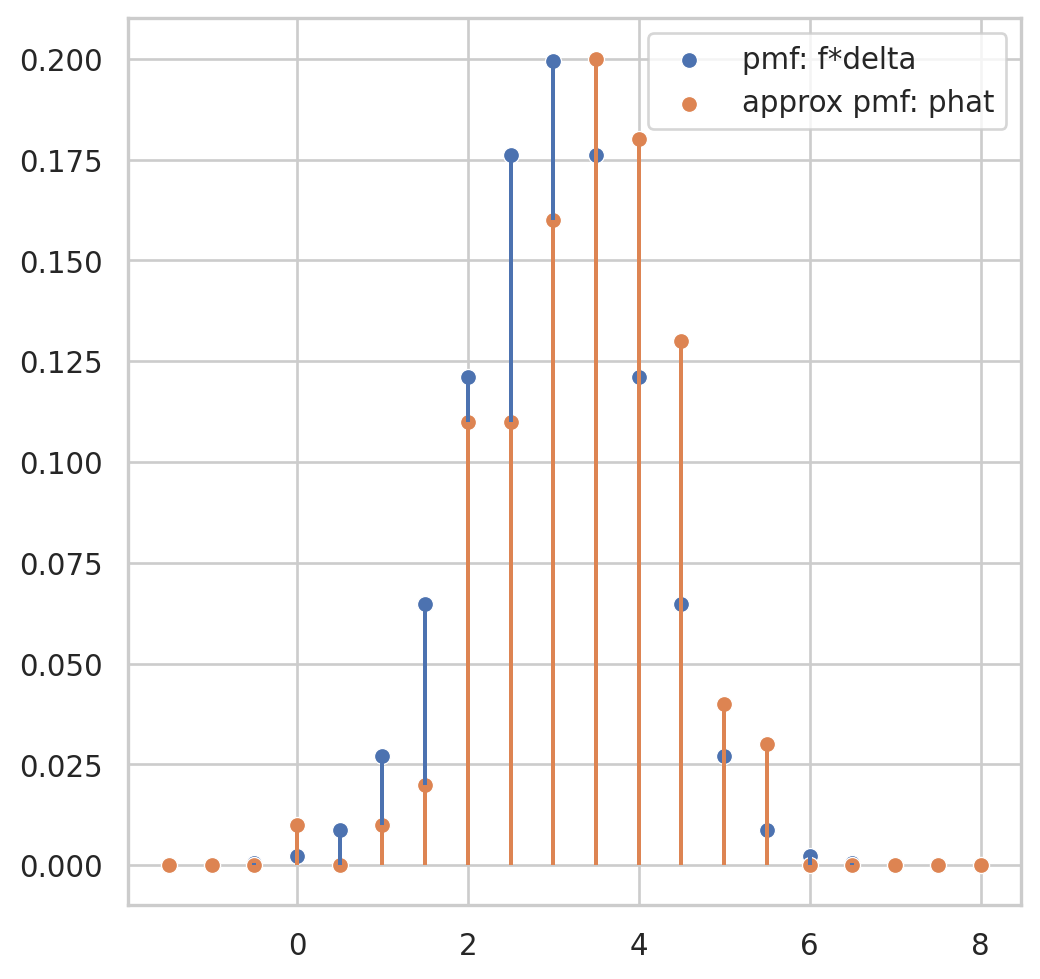
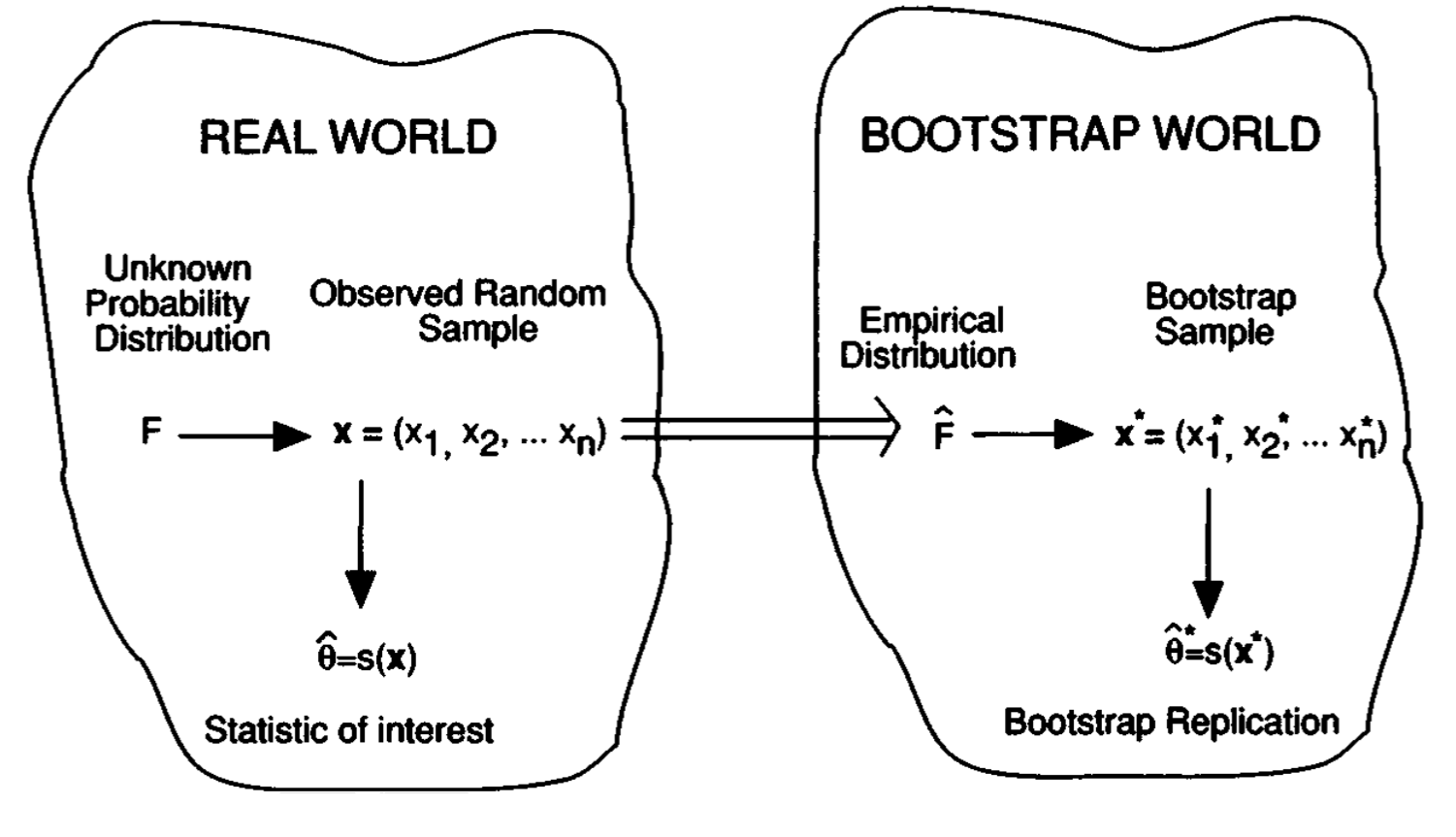
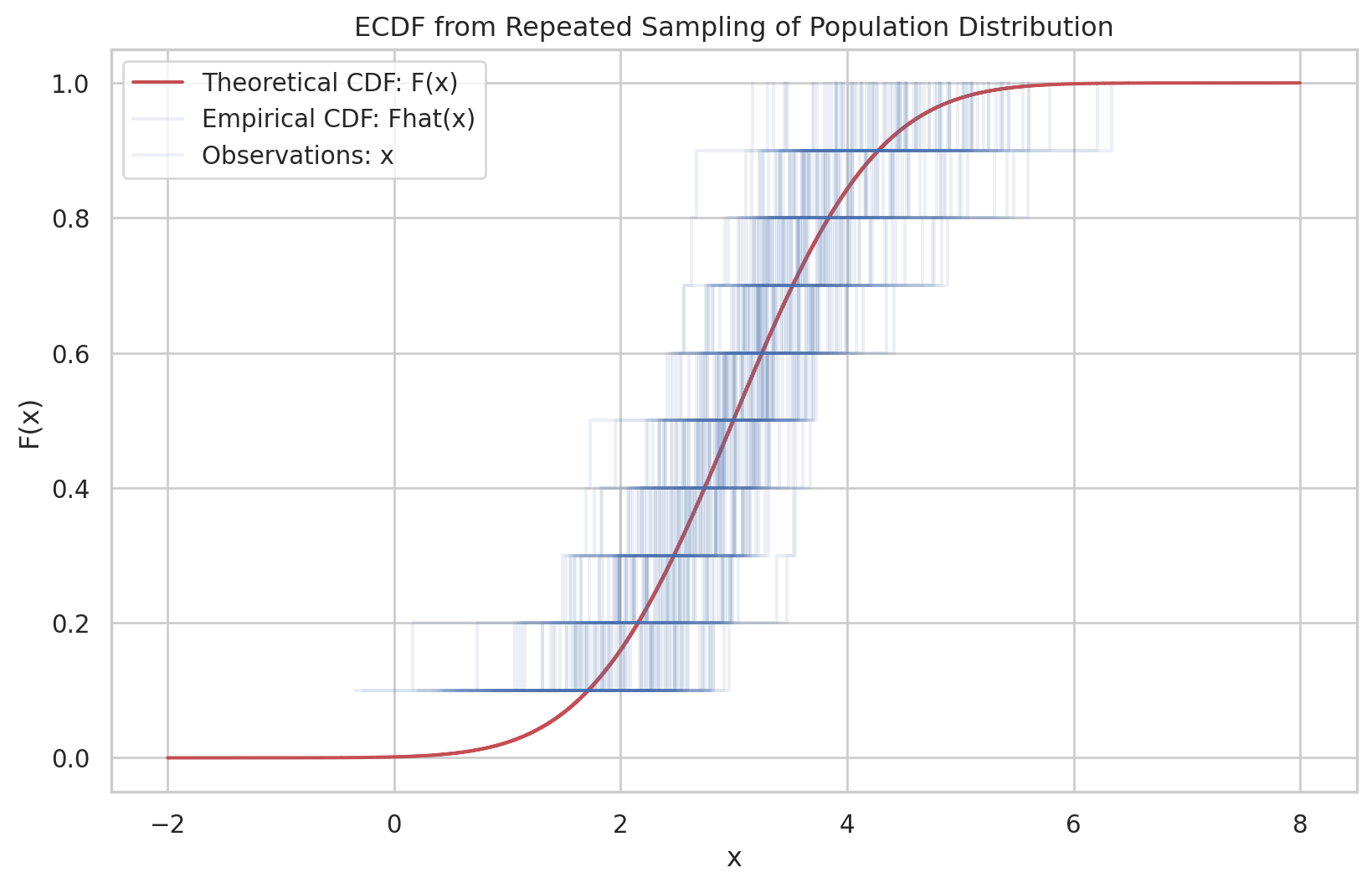
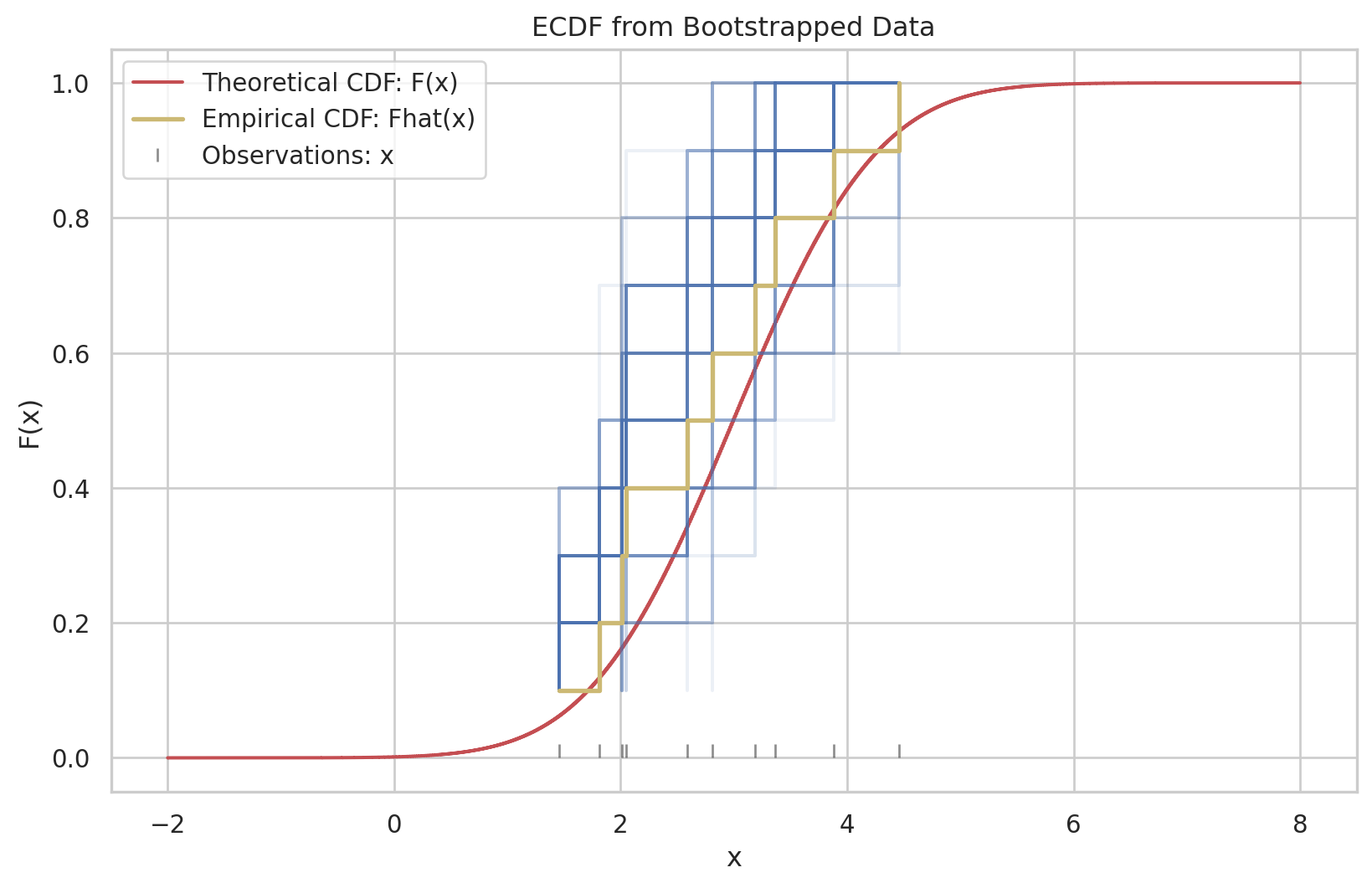
Having observed a random sample of size \(n\) from a probability distribution \(F\),
\[
F \rightarrow\left(x_1, x_2, \cdots, x_n\right)
\]
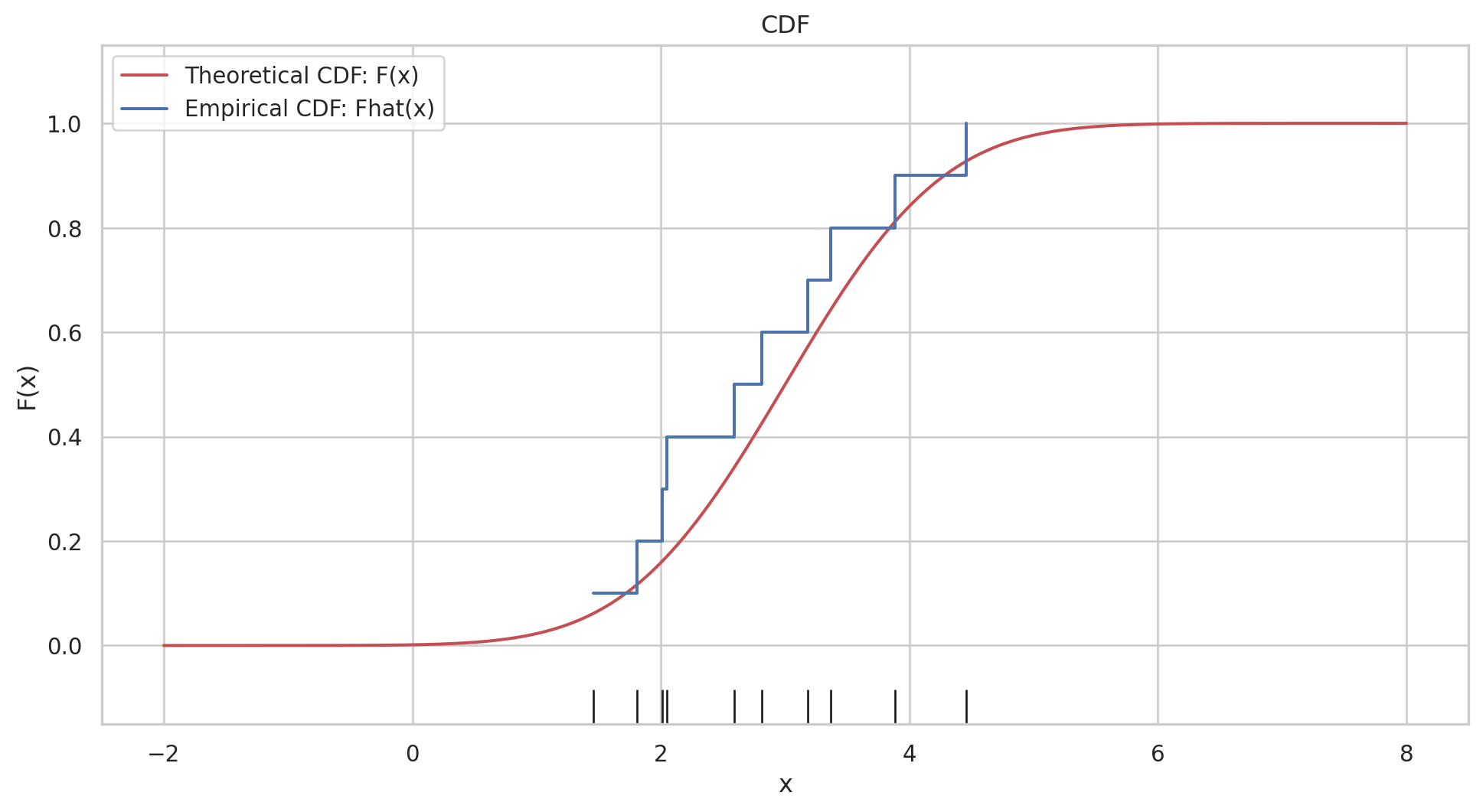
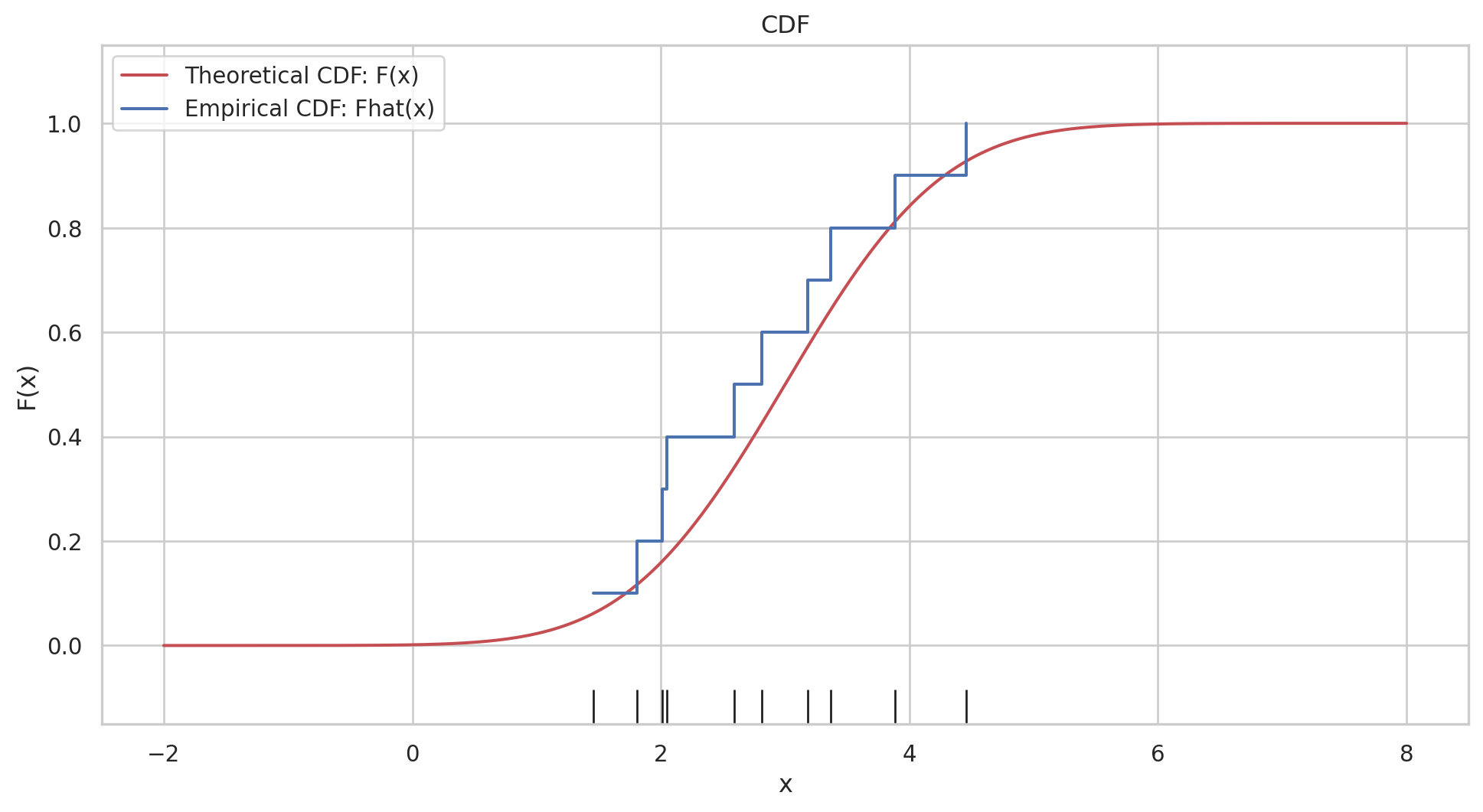
the empirical distribution function \(\hat{F}\) is defined to be the discrete distribution that puts probability \(1 / n\) on each value \(x_i, i=\) \(1,2, \cdots, n\). In other words, \(\hat{F}\) assigns to a set \(A\) in the sample space of \(x\) its empirical probability
\[
\widehat{\operatorname{Prob}}\{A\}=\#\left\{x_i \in A\right\} / n
\]
the proportion of the observed sample \(\mathbf{x}=\left(x_1, x_2, \cdots, x_n\right)\) occurring in \(A\). We will also write \(\operatorname{Prob}_{\hat{F}}\{A\}\) to indicate \(\widehat{\operatorname{Prob}}\{A\}\). The hat symbol ” \(\wedge\) ” always indicates quantities calculated from the observed data.